# CRDS Scan Analyzer
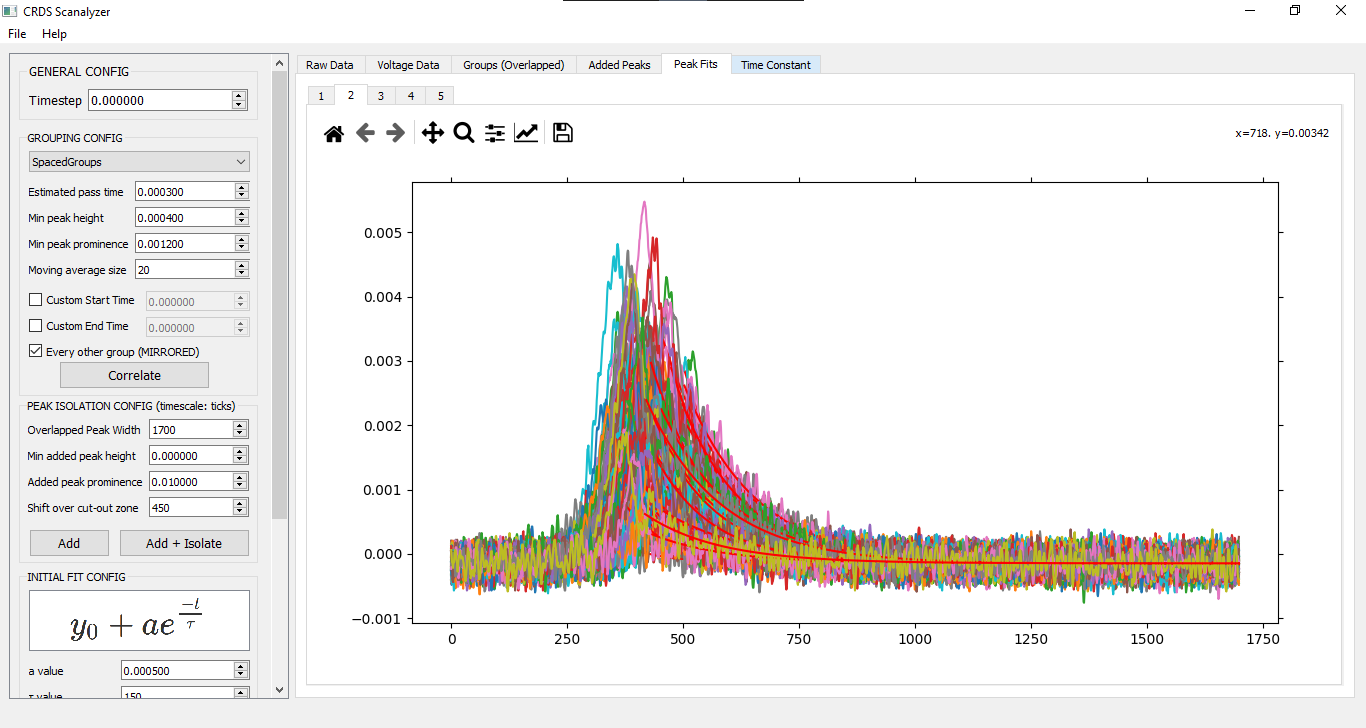 ## Basics
- Designed for usage with **Mid-IR laser comb** scan output (time scale, not frequency)
- Expected data column format: `
## Basics
- Designed for usage with **Mid-IR laser comb** scan output (time scale, not frequency)
- Expected data column format: ` ## Basics
- Designed for usage with **Mid-IR laser comb** scan output (time scale, not frequency)
- Expected data column format: `
## Basics
- Designed for usage with **Mid-IR laser comb** scan output (time scale, not frequency)
- Expected data column format: `